Chapter 3: Midnight Sun
Phase 1 is over :) ! We are half way through the journey. Great learning experience so far. Let’s find out what I accomplished during the previous 2 weeks (since I believe you have been following me from the beginning ;)
Getting straight to the point, most of the time was spent on fixing bugs of the Profiler class and other Pull requests regarding documentation and gallery example. A new gallery example was added to demonstrate the working of SpecDatabase and init_database to help user to store all Spectrums in the form of a .spec file and all input parameters in a csv file under a folder. The same folder can be used to retrieve all Spectrums thus saving a lot of time and also no need to recompute all spectrums, so quite a handy feature. Radis has plot_cond function to plot a 2D heat map based on the parameters in csv file for all spectrums. Creates some good looking and informative plots :)
-> Gallery Example
Back to the analysis part; for LDM we expected:
time(LDM_fft) ~ c2*N_lines + c3*(N_G*N_L + 1)*N_v*log(N_v) (where N_v = Spectral Points)
time(LDM_voigt) ~ c2*N_lines + c3'*(N_G*N_L + 1)*N_truncation*log(N_truncation) (where N_truncation = broadening width / wstep)For Legacy method I was able to prove that Calculation Time is independent of Spectral Range if we keep the Nlines and wstep constant but same is not for LDM voigt.
A straight up comparison between Legacy and LDM voigt for NO keeping Nlines and wstep constant and varying the Spectral range:
Link
Here also for None optimization we are getting constant time for different spectral range but a linear dependency for LDM Voigt which will fail the assumption of
t_LDM_voigt ~ c2*N_lines + c3'*(N_G*N_L + 1)*N_truncation *log(N_truncation )
but rather t_LDM_voigt ~ c2*N_lines + c3*(N_G*N_L + 1)*N_v*log(N_v)A New Discovery
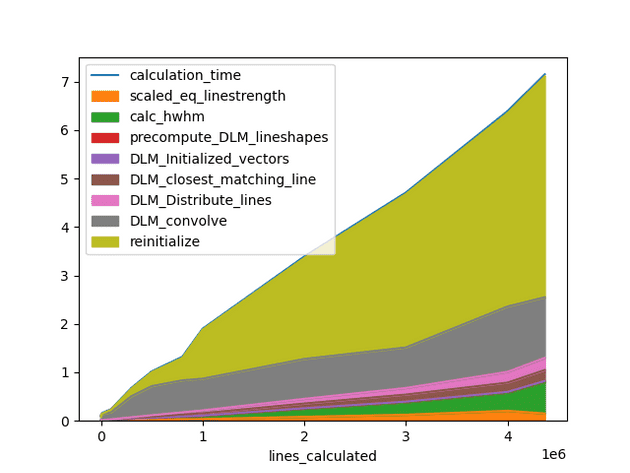
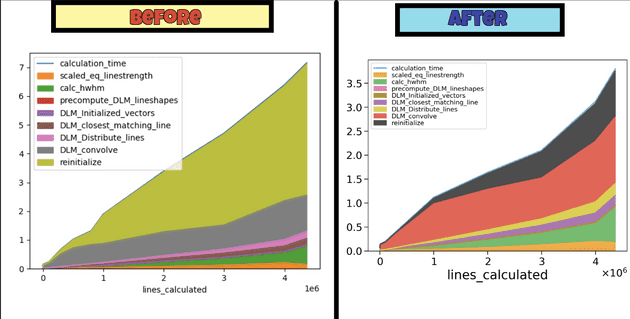
On generating spectrum for millions of lines, one unique observation was seen. The bottleneck step was no longer taking the most time. Max time was spent upon an unknown process. Upon deep analysis it was found a part of code was using sys.getsizeof() to get the size of dataframe, and when the dataframe consisited of object type columns with millions of lines, most of the time was spent on this step only.
We replaced it with memory_usage(deep=False) with a different threshold which made computation almost 2x faster.
So phase 1 is over,and phase 2 is going to begin which will mainly focus on optimizing the the existing LDM method with appropriate truncation and other possible areas!
See you on the other side of the sea ;)